
Recently, a groundbreaking research study was published in the internationally renowned journal *Signal Transduction and Targeted Therapy* (IF = 52.7) by a team led by Academician Bian Xiuwu from the Army Medical University (Director of the Jinfeng Laboratory), in collaboration with the team led by Leng Ling from Peking Union Medical College Hospital and the team led by Zhu Yunping from the National Protein Science Center (Beijing): The world’s first proteomic map covering 13 anatomical regions of the human brain has been successfully constructed. This study not only reveals the specific protein expression patterns in different regions of the brain but also proposes a novel three-module functional framework. This framework provides an important molecular basis for understanding the mechanisms of higher cognitive functions in the brain, as well as the pathogenesis of neurological diseases and the development of targeted diagnostic and therapeutic approaches.

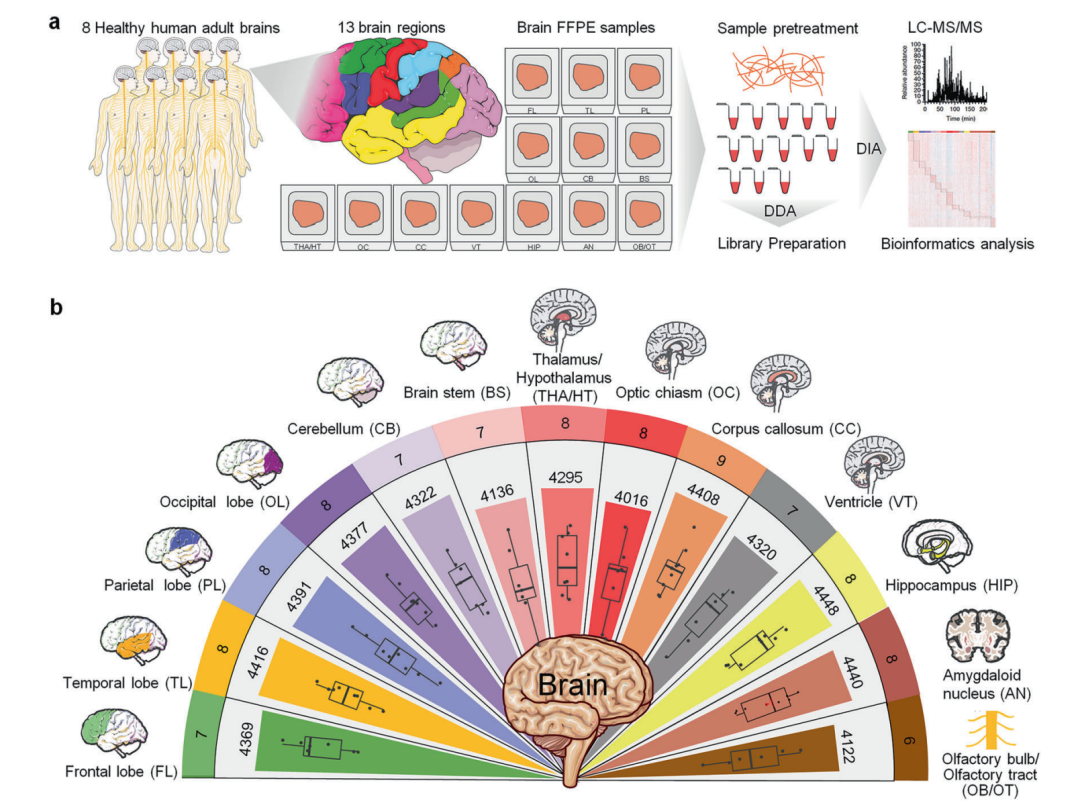
▲Quantitative proteomic analysis of region-specific protein markers in human brain tissue
1. Filling the gaps in research on the brain proteome
Although previous transcriptomic studies have begun to reveal functional differences between brain regions, it remains unclear how the region-specific distribution of proteins, as the direct executors of biological activities, is related to their functions. In this study, deep proteomic analysis was conducted on 13 brain regions from eight cadaver donors, including the four cerebral lobes, the limbic system, and the brainstem. A total of 4,660 proteins were identified, and it was found that 85% to 95% of these proteins were present in all brain regions, although their expression levels exhibited significant regional specificity.
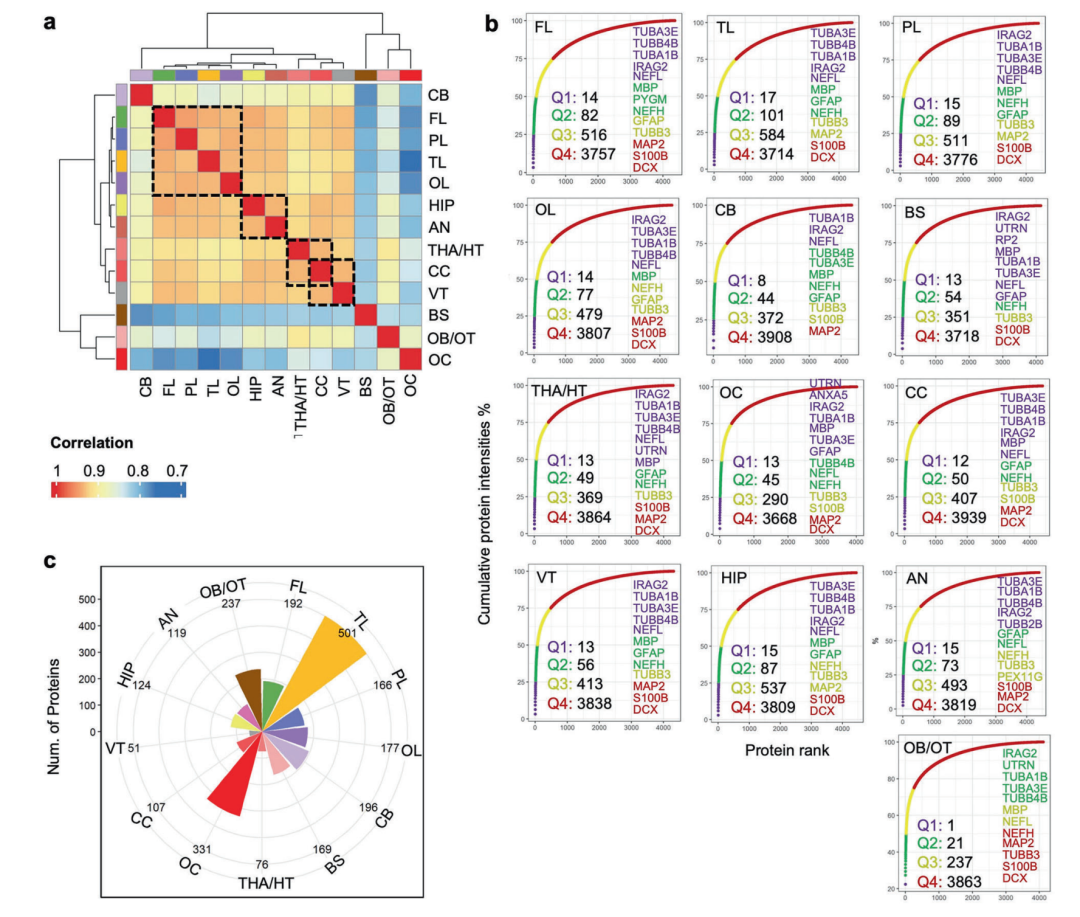
▲Analysis of the differences in proteins with high expression specific to brain regions and their association with brain functional characteristics
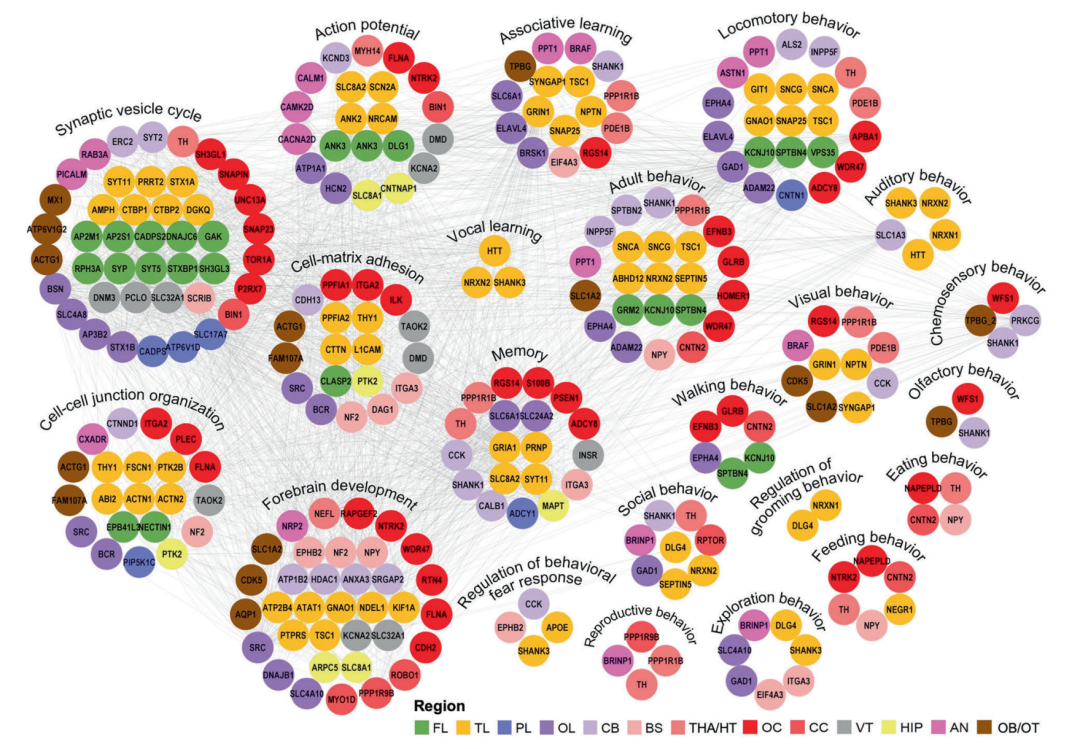
▲Brain region-specific high-expression protein interaction networks constructed based on the STRING database
2. Propose an innovative three-module functional framework
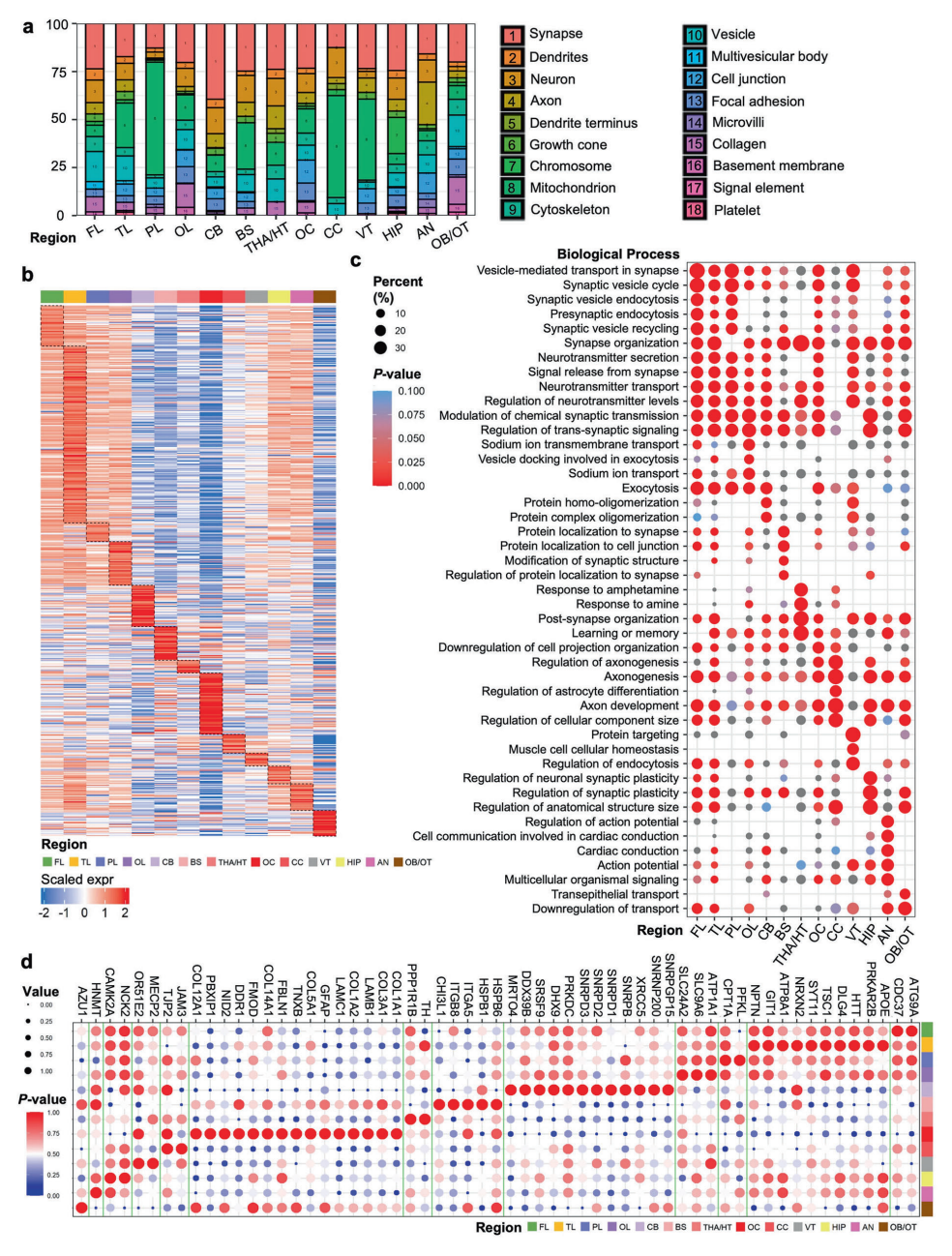
▲Analysis of cellular components in brain regions and functional characterization of neuron-specific overexpressed proteins
Based on the correlation between protein expression similarities and anatomical structures, the research team proposed a completely new organizational module for brain function:
• Cortical integration modules (frontal lobe, temporal lobe, parietal lobe, occipital lobe): Responsible for advanced cognitive processes and information integration ;
• Edge system relay networks (amygdala, hippocampus, thalamus/hypothalamus): Regulating emotions and memory ;
• Midline adjustment axis (thalamus/hypothalamus, corpus callosum, ventricles, optic chiasm): For the first time, it has been demonstrated to play a central role in neural development, inter-regional signal transmission, and structural homeostasis.
The study suggests that the midline regulatory axis may contribute to the overall stability of the brain through efficient synaptic function, metabolic processes, and the integrity of the extracellular matrix. This finding challenges the traditional view of it as a “passive structure.”
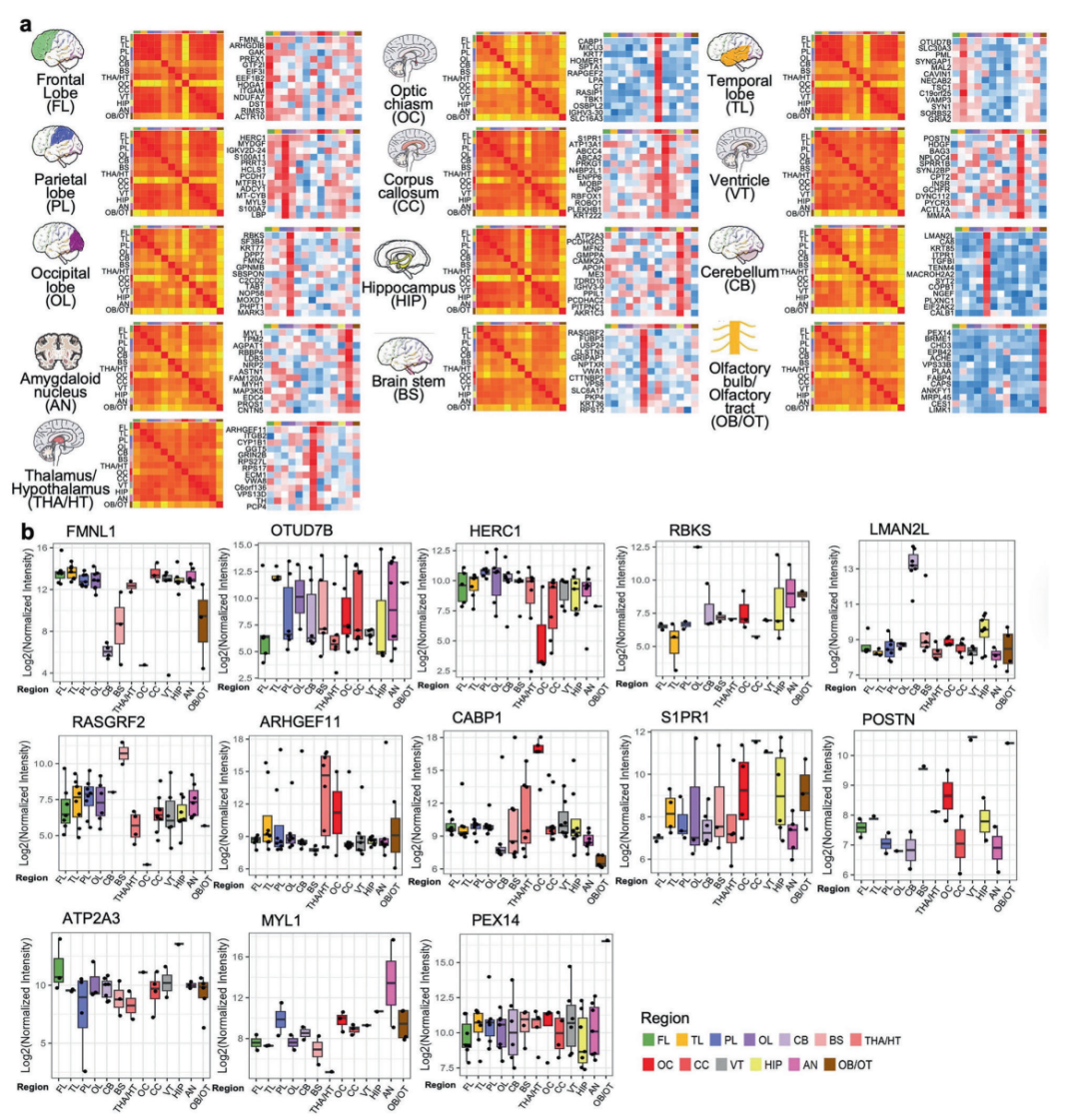

▲Comparative analysis of highly expressed protein profiles in different brain regions and their cross-regional expression patterns
3. The temporal lobe has the highest protein complexity, and its ventricular functions are surprisingly active
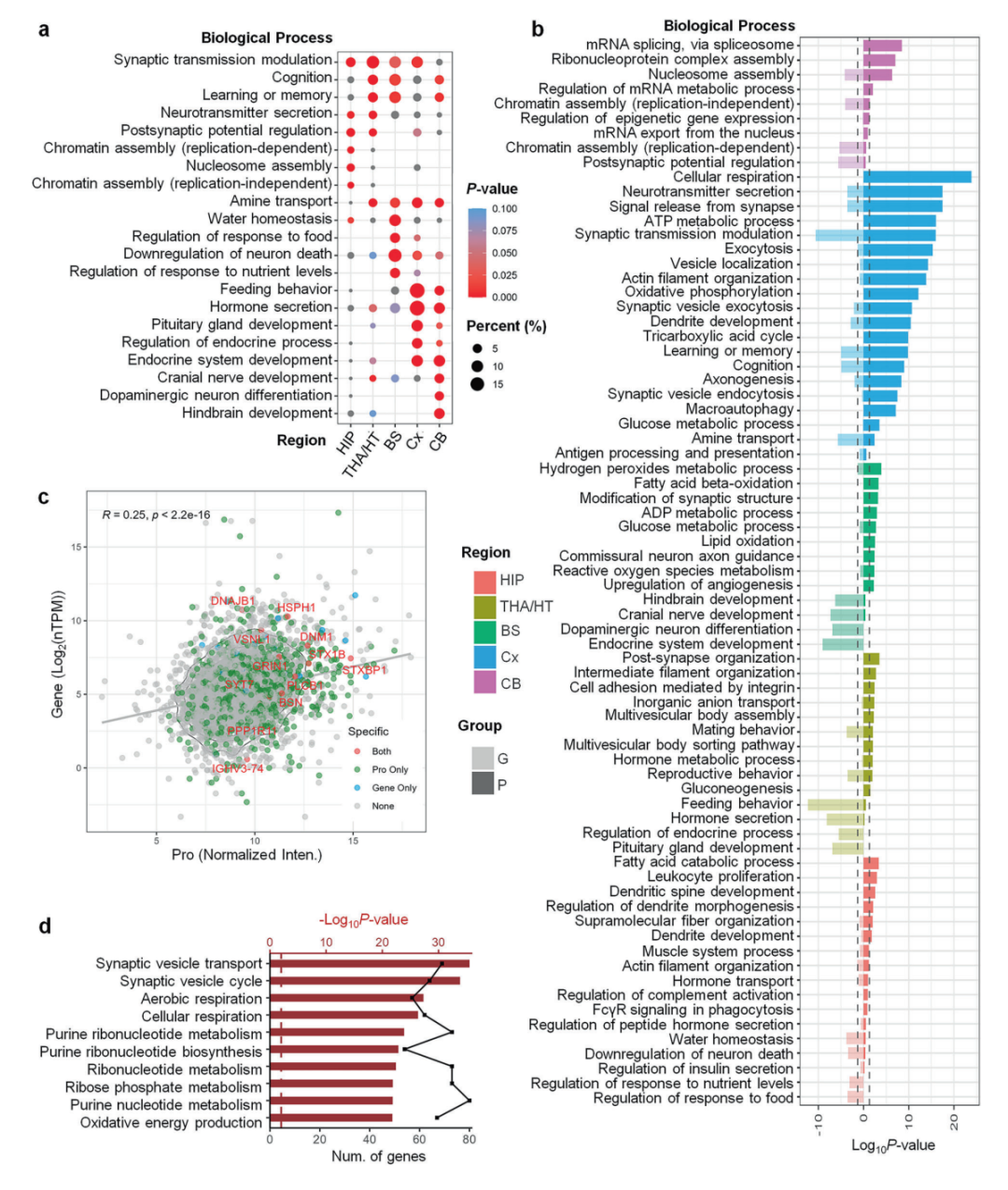
▲Comparison of results from human brain pathway analysis using proteomics and transcriptomics
The research results indicate that the temporal lobe contains 501 region-specific proteins that are highly expressed, the highest number of any brain region, highlighting its central role in auditory and visual integration as well as memory processing. Regions of the ventricles, which are traditionally considered to have single functions, have been shown to exhibit high expression of proteins related to neural development, glial cell differentiation, and energy metabolism, suggesting that they may play an active role in regulating brain homeostasis.
4. Research Significance and Future Prospects
This study found that, by integrating proteomic and transcriptomic data, protein levels are a better reflection of the actual functional state of brain regions. The team further verified several potential biomarkers: for instance, the high expression of the POSTN protein in the ventricles may be associated with congenital hydrocephalus, while the S1PR1 protein in the corpus callosum could provide diagnostic clues for multiple sclerosis. These findings open up new avenues for the early diagnosis and targeted treatment of neurological disorders. This study not only provides a high-resolution map of the human brain’s proteome but also redefines the molecular basis of brain functional modules. In the future, by combining multi-omics techniques with clinical data, it is expected that the molecular mechanisms underlying brain-specific damage in diseases such as Alzheimer’s and autism will be elucidated.
The first author of this paper is Associate Chief Physician Zhang Peipei, a postdoctoral researcher at both the Jin Feng Laboratory and the Mobile Postdoctoral Station of the School of Basic Medicine at Army Medical University (Jin Feng Laboratory/Army Medical University/Shanghai Jiao Tong University). The co-first authors include Li Mansheng, an associate researcher at the National Protein Science Center <北京> ) and Zhou Jia (a doctoral student in the eight-year clinical program at Peking Union Medical College Hospital, Chinese Academy of Medical Sciences). Leng Ling (Peking Union Medical College Hospital, Chinese Academy of Medical Sciences), Bian Xiuwu (Army Medical University), and Zhu Yunping (National Protein Science Center) <北京> ) are co-corresponding authors.
Click “Read the original text” in the lower left corner to access the link to the original article.